Identifying the progression of proteins underlying Parkinson's Disease
PTI is applying its high-depth and single-cell proteomics technologies to generate an unprecedented view of protein dysfunctions in Parkinson's disease, with the goal of uncovering causal mechanisms and new therapeutic targets.
Diagnosing and mechanistically understanding early-stage Parkinson's disease (PD) requires deep, high-throughput proteomic analysis of disease-specific protein modifications. PTI’s technologies are enabling such analysis at unprecedented scale and resolution: For the first time, we can analyze the proteomes of thousands of single cells from PD brains and reconstruct disease progression at single-cell resolution to guide therapeutic development.
Parkinson's disease is the second most common neurodegenerative disorder, affecting more than 10 million people worldwide (Parkinson's Foundation). Its defining hallmark is the misfolding and aggregation of alpha-synuclein into Lewy bodies. However, the protein dysfunctions underlying PD extend beyond this single protein. Understanding what goes wrong across thousands of proteins in individual cell types and at the earliest disease stages requires technologies capable of measuring proteomes at a scale and resolution not previously possible. PTI was built to close this gap.
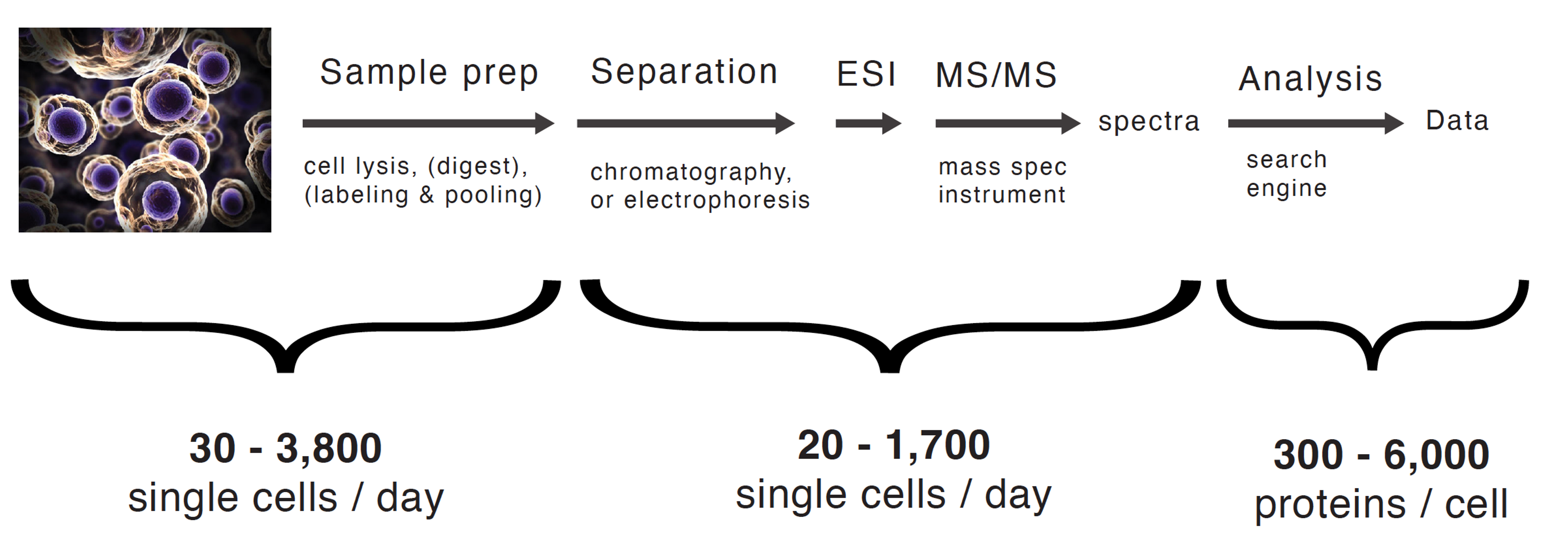
Figure 1. PTI's single-cell proteomics workflow, enabling quantification of 300–6,000 proteins per cell across up to 3,800 single cells per day. PTI & Slavov Lab; Leduc et al., Nature Protocols, 2024.
A key advance in PTI's ability is to directly measure alpha-synuclein and other PD-relevant proteins at single-cell resolution and analyze how they co-vary across individual neurons and glia, a capability not previously demonstrated. PTI's platform enables quantification of hundreds to thousands of proteins per cell, revealing changes in specific cell populations (Derks et al., Nat. Methods, 2023; Huffman et al., Nat. Methods, 2023).
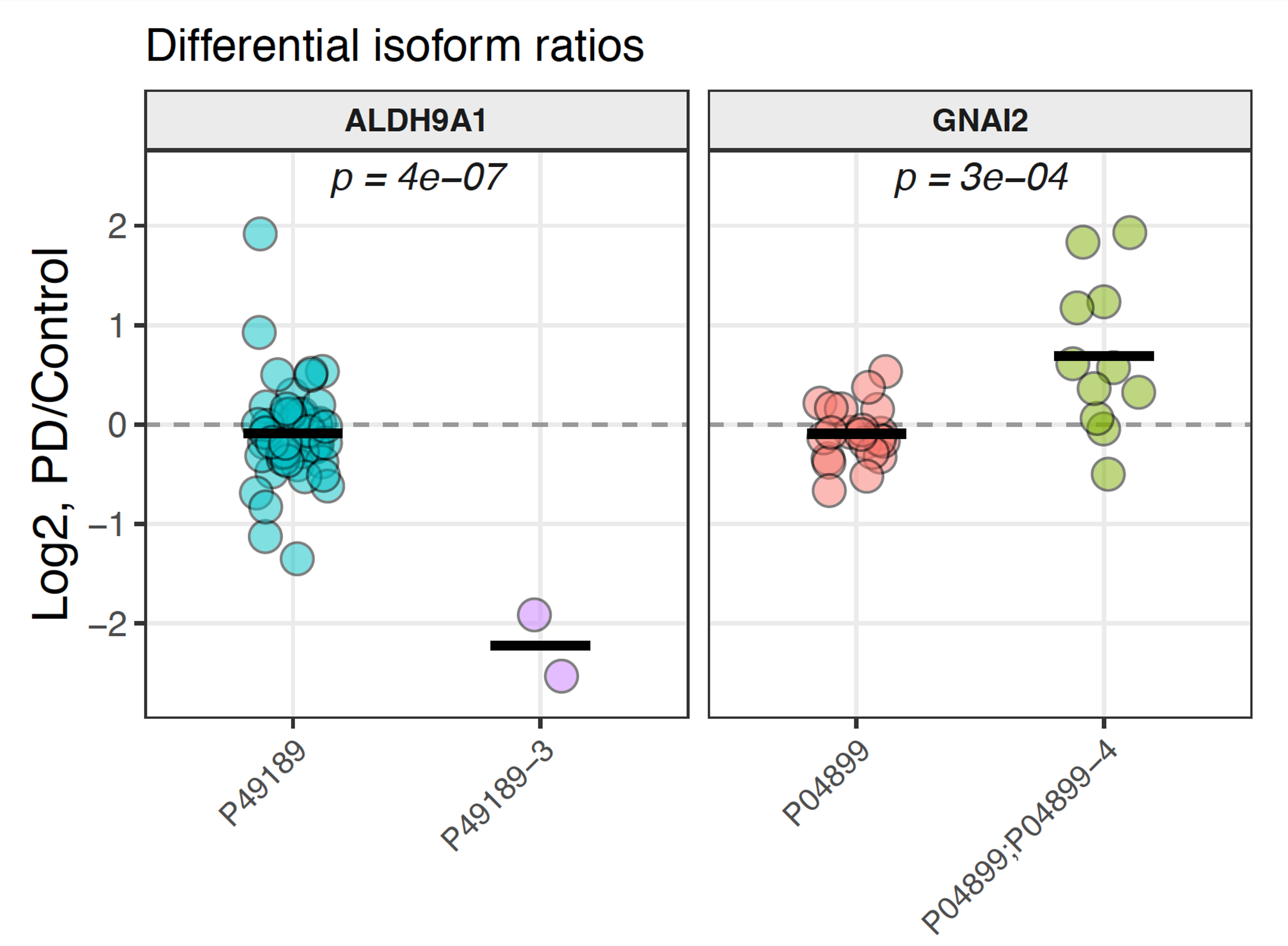
Figure 2. Differential isoform ratios for ALDH9A1 and GNAI2, PD vs. control. PTI & Slavov Lab, PDRx 2026.
PTI's large-scale peptide-level measurements also resolve proteins in their distinct molecular forms across patient samples. Proteins such as ALDH9A1 and GNAI2 show isoform-ratio differences between PD and controls invisible to standard abundance measurements (Fig. 2), and alpha-synuclein peptide profiling reveals patient-specific patterns that bulk proteomics averages out (Fig. 3). PTI additionally detects modifications such as differential phosphorylation at the level of the proteome, identifying molecular casualties of early pathology previously inaccessible (Fig. 4). Across all samples, PTI's plexDIA platform quantified 10,426 proteins per sample and identified 259 significantly differentially abundant proteins (q < 0.05), including TH and DDC.
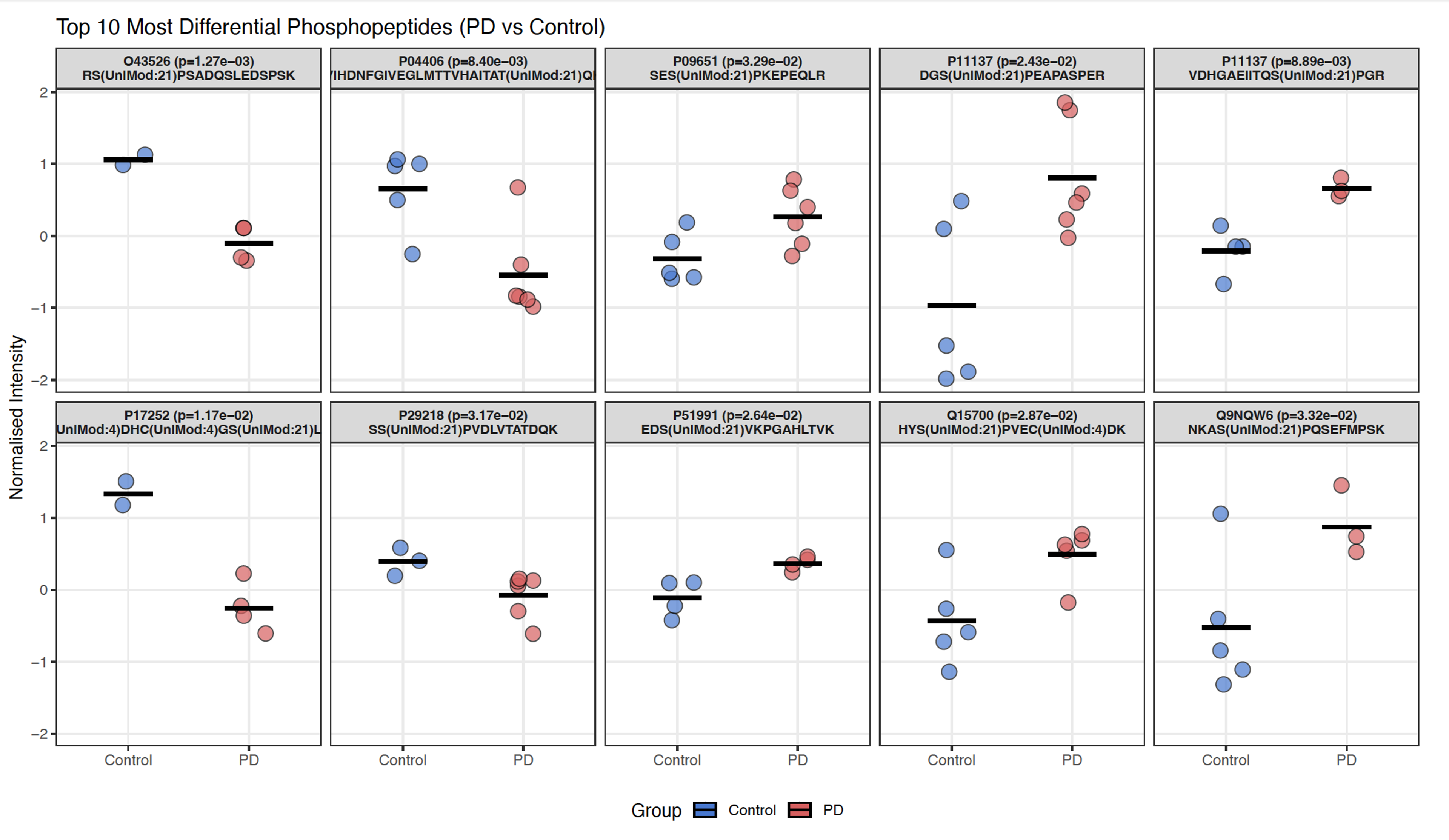
Figure 4. Top differential phosphopeptides, PD vs. control. PTI & Slavov Lab, PDRx 2026.
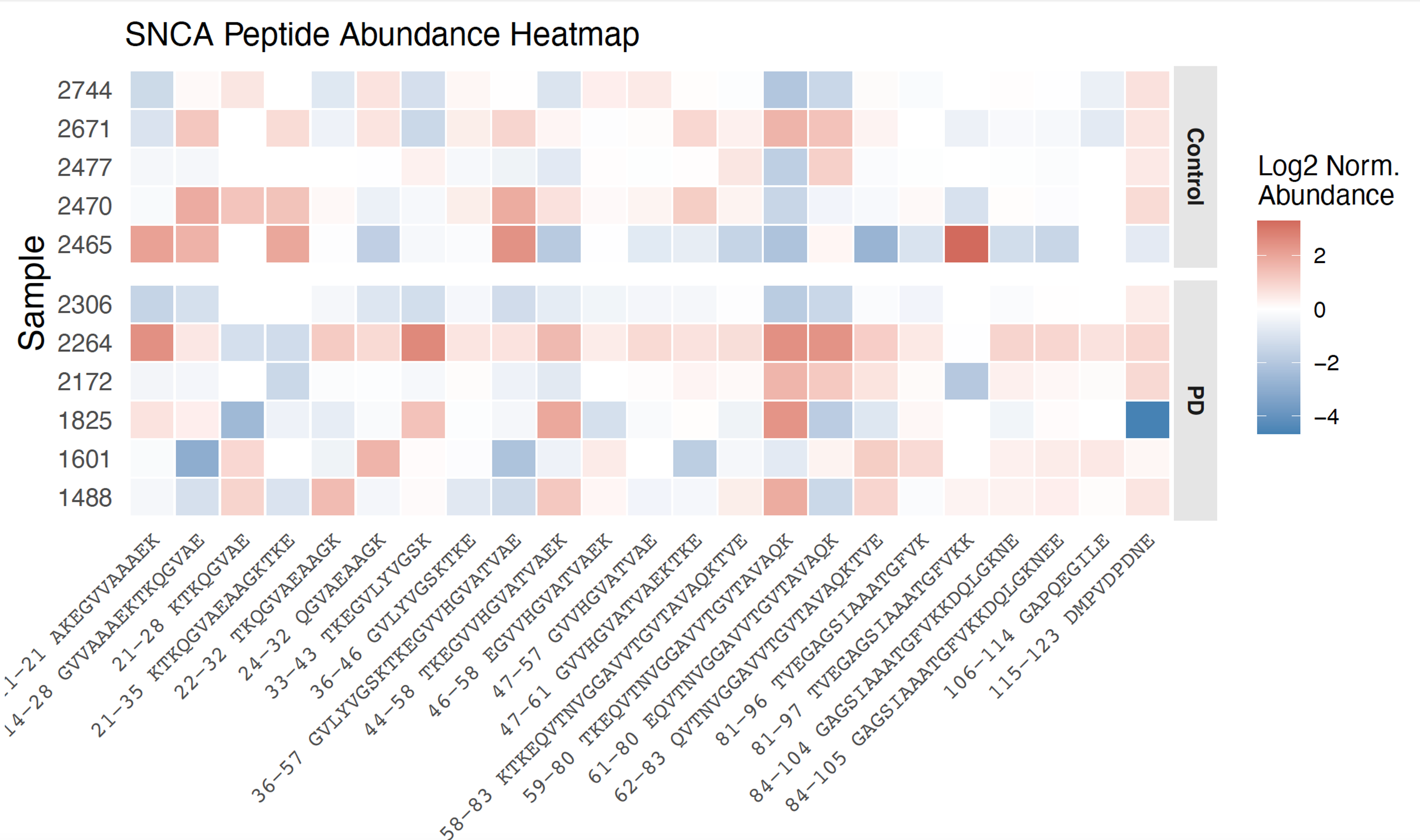
Figure 3. SNCA peptide abundance heatmap, PD vs. control. PTI & Slavov Lab, PDRx 2026.
By openly sharing all data and resources, PTI aims to shift Parkinson's research from late-stage bulk characterization toward early detection and mechanistic understanding, where therapeutic impact is greatest.