We aim to increase the throughput of mass spectrometry proteomics 100-1000x
Technology
We have already demonstrated a proof-of-principle through the development of plexDIA. That endeavor increased the sensitivity of low-input samples using commercially available reagents. We now aim to scale the throughput by 100-1000 fold by synergistically optimizing mass spectrometry methods, creation of novel barcodes, and implementing deep learning analysis. This multiplicative increase in throughput arises from the simultaneous parallelization of both samples (single cells per unit time) and the depth of proteome coverage (peptides quantified per sample). This inspires the name of this FRO, Parallel Squared.
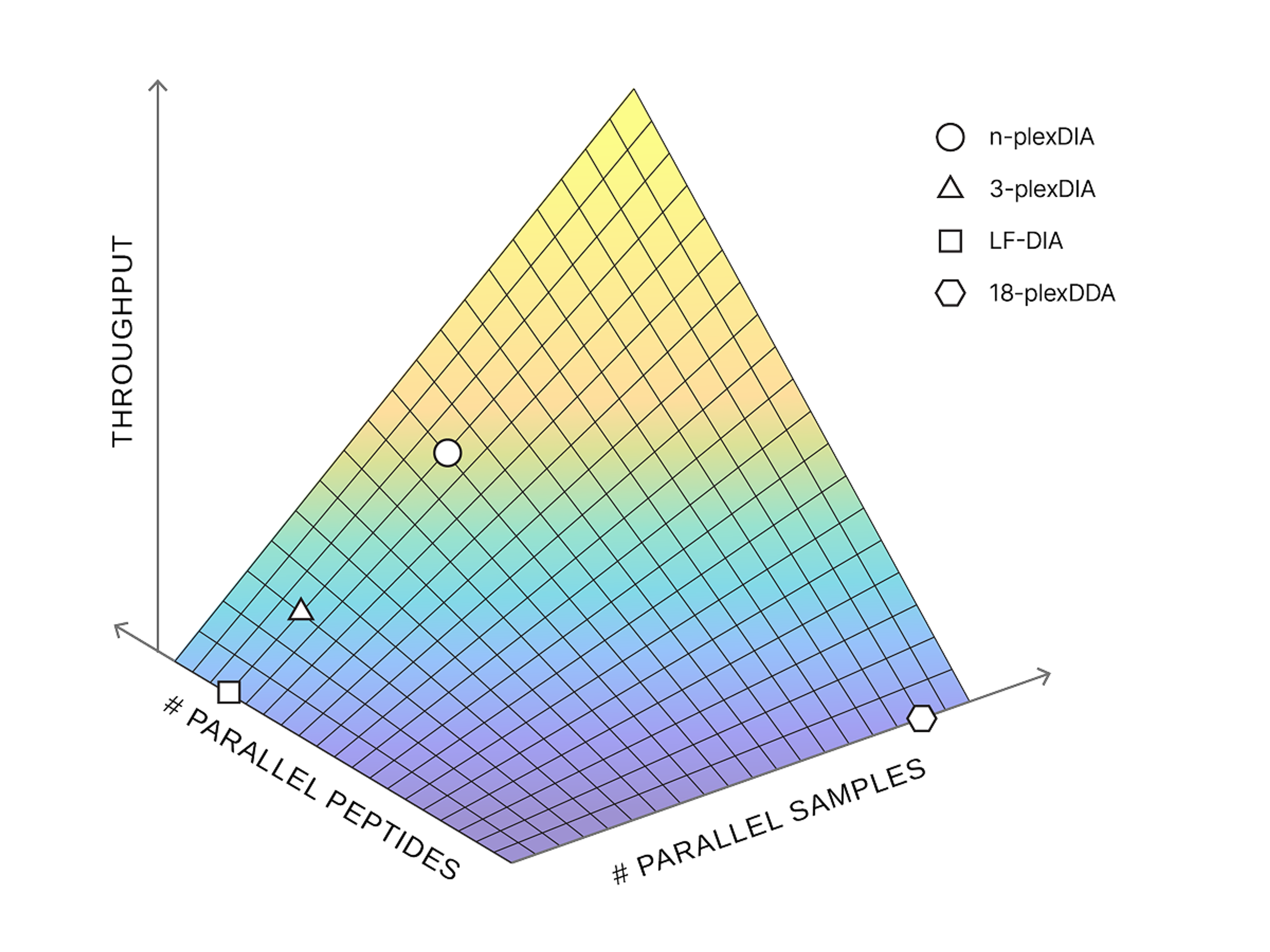
plexDIA
plexDIA is a multiplexing method that triples proteomics throughput from tiny samples without losing accuracy, enabling fast, high-coverage analysis—even at the single-cell level.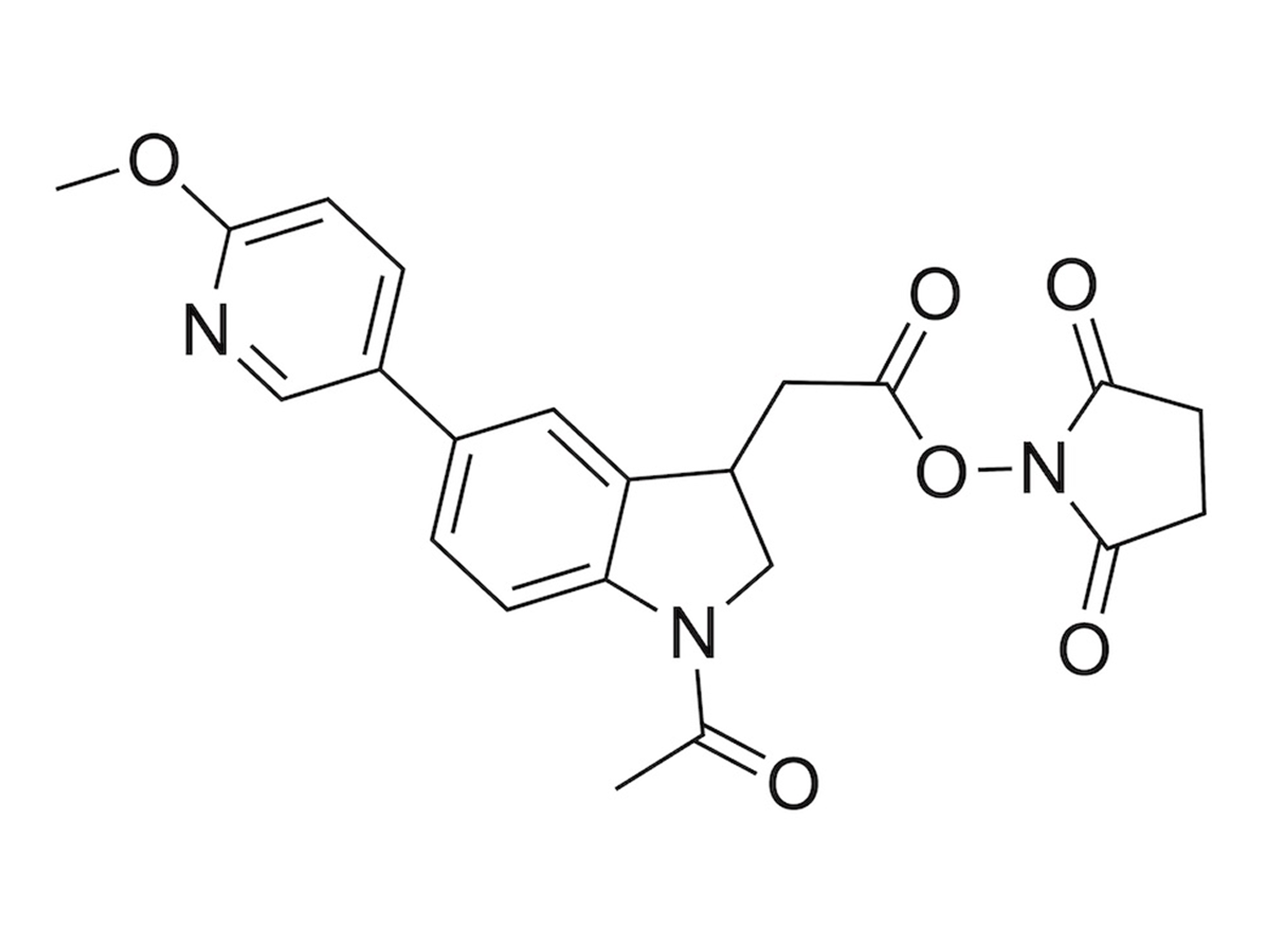
PSMtags
A new mass tag enabling 9-plexDIA with better sequencing and multiplexing, allowing high-throughput proteomics of over 1,000 samples per day.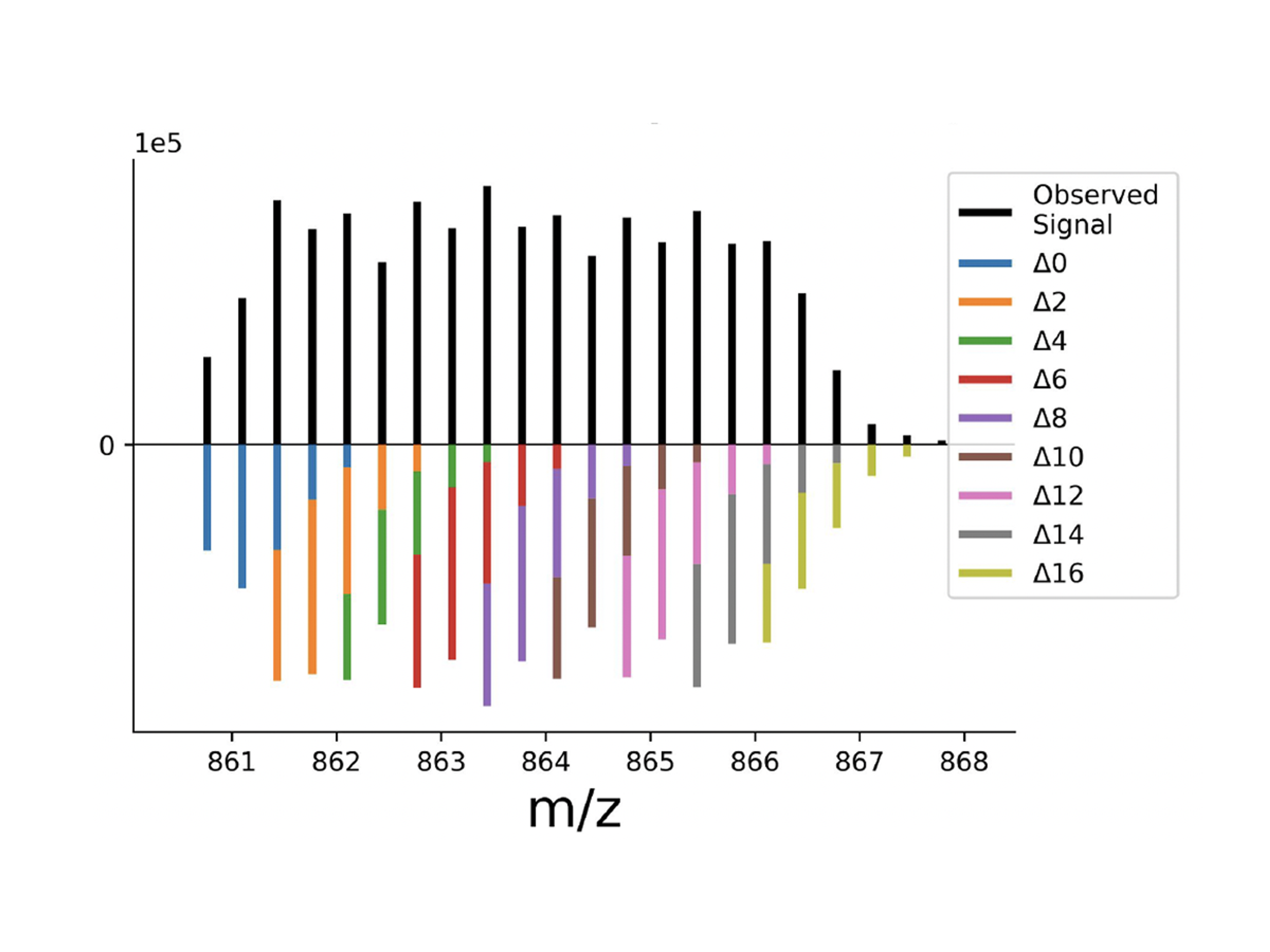
JMod
JMod is an open-source platform that models overlapping mass spectra, enabling accurate sequence identification and quantification from highly multiplexed data.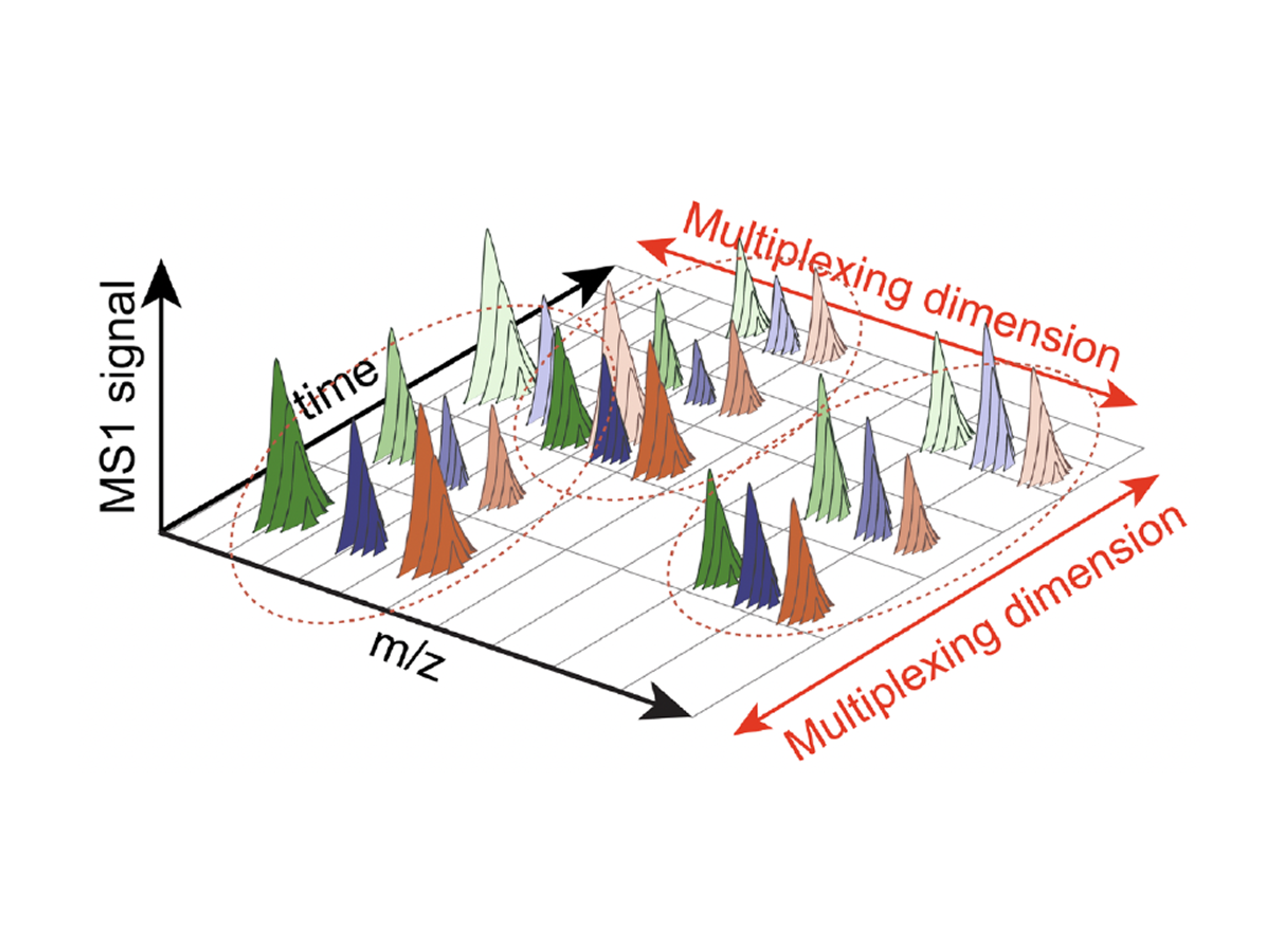
timePlex
timePlex acheives sample separation by staggering them in the time domain, providing a multiplexing strategy which enables combinatorial scaling to over 500 samples per day.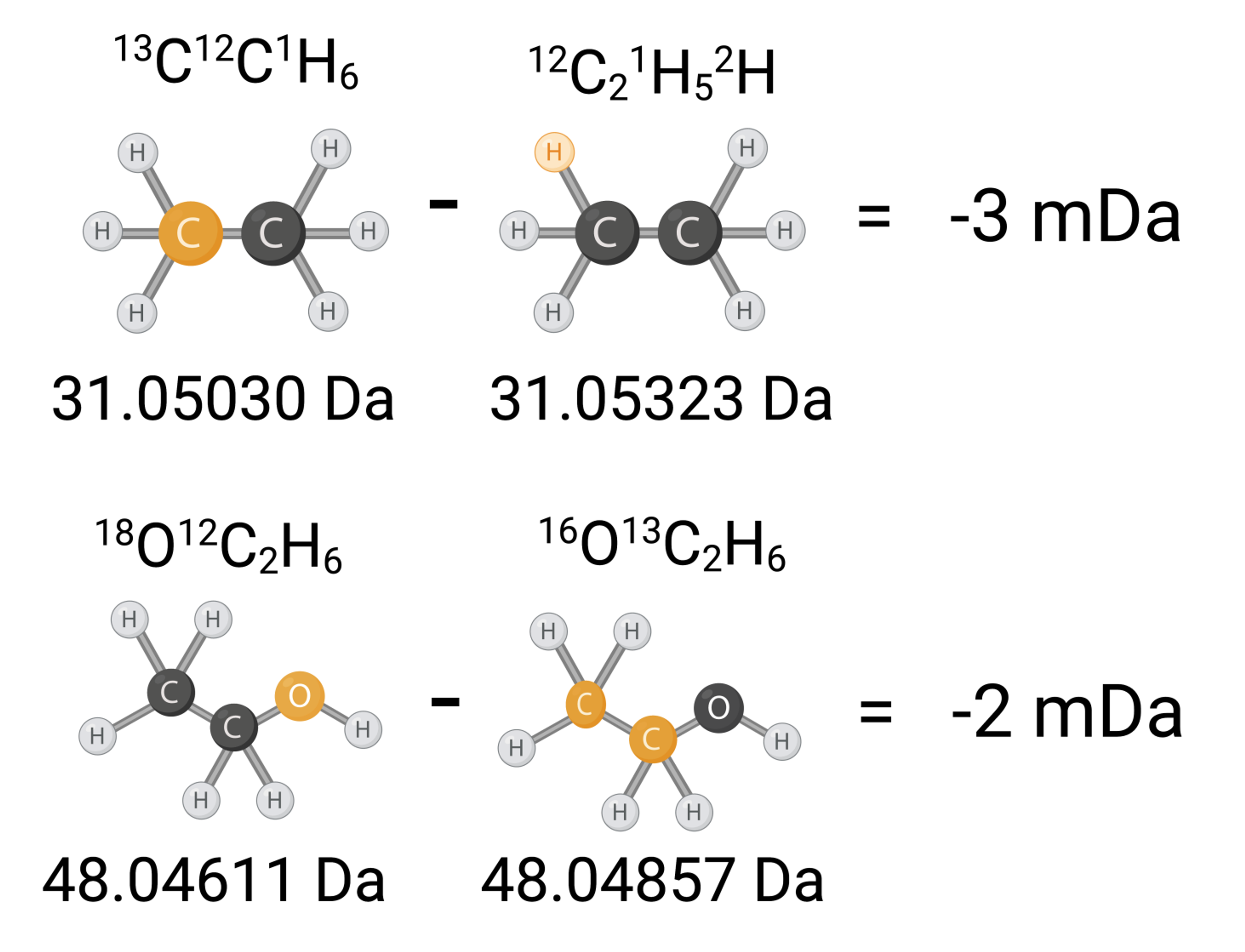
Tag Design
A framework for designing mass tags using the differential mass defect to demonstrate that plex sizes in the hundreds are achievable with molecules comparable in size to existing commercial tags.Neurodegenerative Diseases
Neurodegenerative diseases such as Parkinson's and Alzheimer's are driven by protein dysfunctions that remain poorly understood because the technologies needed to measure them at sufficient resolution have not existed. PTI is streamlining the process to close this gap. By quantifying thousands of proteins, their isoforms, and modifications in individual neurons and glia from patient tissue, PTI is resolving the causal drivers of neurodegeneration to guide therapeutic intervention at the earliest and most effective stages.
Alzheimer's Disease
To better understand Alzheimer’s, our approach analyzes protein data at two scales—brain voxels to track spatial disease progression, and single cells to reveal how diverse cell types are differently affected.